Biochemistry and Molecular Biology
Biochemistry and Molecular Biology
Weiskotten Hall
Rm. 4265
766 Irving Avenue
Syracuse, NY 13210
Google Maps & Directions
Phone: 315 464-5127
Fax: 315 464-8750
Name: Stewart Loh, PhD, Interim Chair
Email: [email protected]
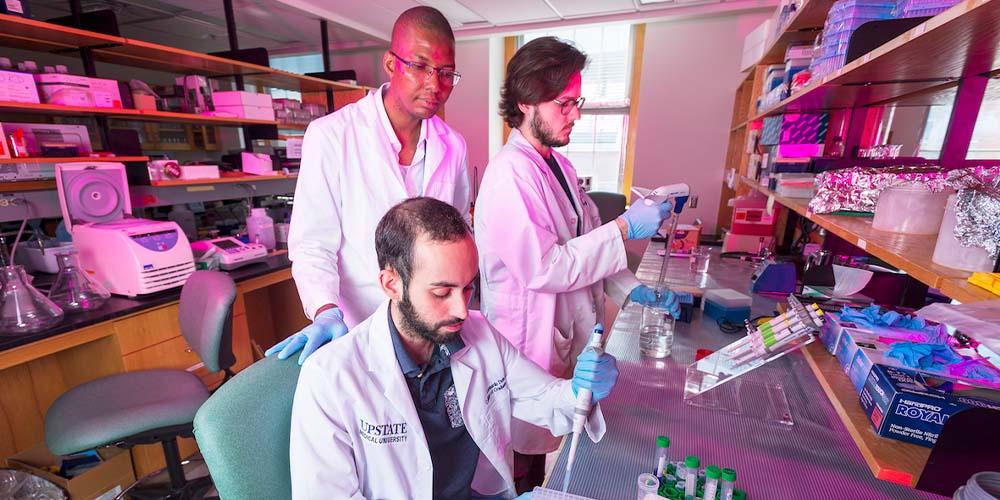
The Department of Biochemistry & Molecular Biology offers a highly collaborative community in which researchers employ state-of-the-art equipment, facilities, and approaches to investigate molecular mechanisms that form the basis of life. The fields of biochemistry and molecular biology have been transformed by recent innovations in genomics, structural biology, computational tools, and instrumentation technology. The integration of these advancements creates unprecedented opportunities for scientific research and breakthroughs in the understanding and treatment of disease. Our faculty work closely with postdocs and students to provide a vibrant mentoring and training experience for young scientists. Weiskotten Hall
Rm. 4265
766 Irving Avenue
Syracuse, NY 13210
Google Maps & Directions
Phone: 315 464-5127
Fax: 315 464-8750
Name: Stewart Loh, PhD, Interim Chair
Email: [email protected]
WELCOME

Stewart Loh, PhD
Interim Chair and Professor
In the department of Biochemistry & Molecular Biology, our goals are to: (1) uncover fundamental mechanisms of living systems, from molecular structures and interactions to cellular processes to human health and disease; (2) train the next generation of young scientists. Read More...
Research Highlight
- Oxr1 - a novel regulator of V-ATPase
The vacuolar ATPase (V-ATPase) is a highly conserved rotary motor proton pump that plays an essential role in cellular housekeeping functions. Not surprisingly, V-ATPase activity (or loss thereof) has been linked to several disease states including renal tubular acidosis, osteoporosis, neurodegeneration, and cancer. V-ATPase is made of two subcomplexes: a cytosolic V1 that carries out ATP hydrolysis, and a membrane bound Vo that is responsible for proton translocation. The enzyme’s proton pumping function is regulated by a unique process called “reversible disassembly”, wherein V1 dissociates from Vo in a for example nutrient dependent manner. To understand V-ATPase’s role in health and disease, the Wilkens lab studies the structure and mechanism of the enzymes from yeast and human. Read more... - Implication of Organelle pH Regulation by Phosphoinositides through “Zip Codes” in Vacuolar H+-ATPase

The organelles of the secretory and endocytic pathway include the ER, Golgi network, endosomes and lysosomes or, the lysosome like yeast vacuole. All of these organelles maintain a distinct and tight range of pH. The vacuole/lysosome (pH 4/5.5) is the most acidic compartment in eukaryotes, whereas the Golgi is relatively alkaline (pH 6.6). Variation in pH largely affects the function of these organelles. For example, alkaline vacuole/lysosome are deficient in autophagy, Golgi pH regulates its ability to glycosylate proteins and failure to maintain endosomal pH perturbs with its ability to recycle receptors to the Plasma membrane or, the trans-Golgi. Read more... - Structure of Lipid Nanodisc-reconstituted Vacuolar ATPase Proton Channel: Defining the Interaction of Rotor and Stator

The Wilkens lab focuses on the structural characterization of the vacuolar ATPase (or V-ATPase) in order to elucidate its mechanism of activity and regulation. V-ATPase is a ubiquitous eukaryotic rotary proton pump that is foiund in the endomembrane system and serves to acidify a variety of intracellular compartments as well as the extracellular space in higher eukaryotes. V-ATPase is essential in animals; full loss of activity is embryonic lethal and partial loss of function has been associated with various human diseases including osteoporosis, male infertility, sensorineural deafness, diabetes and various cancers. Read more...
- The Smc5/6 complex at the crossroads of DNA replication, repair and recombination

Due to the imperfect “steric gate”, DNA polymerase intrinsically mis-incorporates not only mismatched deoxyribonucleosides monophosphates but also ribonucleoside monophosphates (rNMPs) during DNA replication at a rate of 10−7 and 4 × 10−4, respectively (1). To repair these DNA damages it requires specific recognition and excision proteins to remove the damage and create a single-stranded DNA (ssDNA) gap, followed by the DNA polymerase to fill the gap and the DNA ligase to seal the nick. Read more...
- New class of p53-reactivating compounds provides novel mechanism to treat cancer - August 2015

The "Guardian of the Genome," p53, is a tumor suppressing transcription factor that has long been recognized as perhaps the most important protein in human cancer. Approximately 50% of human cancers harbor mutations in p53, which render the protein inactive and unable to protect the cells from cancerous transformation. Read more...
- Mechanism of Ant1-induced human diseases unraveled by the Chen lab - July 2015

Mitochondria are the powerhouses of the cell. About 90% of the energy that cells need is produced in the form of ATP by the OXPHOS apparatus on the mitochondrial inner membrane. After it's synthesis by the F0F1 ATP-synthase, ATP is exported out of mitochondria via adenine nucleotide translocase (Ant) by exchanging with the cytosolic ADP. Read more...
- Using disease-associated mutations to understand the biochemical regulation of a multi-subunit histone methyltransferase complex - June 2014

Eukaryotic DNA is compacted into chromatin, which must be continually remodeled to allow for DNA processes such as transcription. The basic repeating unit of chromatin, the nucleosome, is composed of an octomer of histone proteins around which 147 base pairs of DNA is wrapped. One way chromatin remodeling is achieved is by posttransitional modification of histones. Read more...




